
Ellen Hsu, PhD
Professor
Physiology and Pharmacology
The central problem in immunology is to understand how an organism distinguishes self from foreign substances. The first process is the acquisition of immunocompetence, the second, tolerance induction; both are mediated through recognition by lymphocytes via cell surface molecules. Our research concerns the mechanisms by which the vast repertoire of antigen receptors (immunoglobulin, T cell receptor) is generated and how these processes evolved.

Figure 1. Sharks belong to the class Chondrichthyes, whose fossils first appear in the early Devonian, 400 million years ago, and are the living representatives of the most primitive jawed vertebrates. They are the earliest chordates to possess TCR, MHC, and Ig genes. Extensive somatic hypermutation in the light chains shows that this mechanism of diversification has played a significant role since the origins of the adaptive immune system (featured article). Photography by William Struhs; taken at the South Carolina Aquarium, Charleston, South Carolina.
The adaptive immune systems of vertebrates are characterized by a primary response mounted specifically against the invading pathogen and, upon re-infection by the same pathogen, a secondary response that is more rapid and of greater magnitude. The specificity and memory are based on the possession of lymphocytes, circulating cells with surface receptors expressing a vast, anticipatory repertoire of ligand-combining sites directed against potential pathogens. Antibody-producing B lymphocytes express immunoglobulin (Ig) on their cell surface while effector and cytotoxic killer T lymphocytes express T cell receptor (TCR). Estimates range about 108-1014 different receptors for humans. Since the human genome carries at the current estimate 25,000 protein-encoding genes, it cannot encode so many antigen-binding receptors in the germline explicitly. In fact, the receptors are somatically generated from component parts by recombination in differentiating lymphocytes during the lifetime of the individual.
Antibody receptor gene segments in tetrapods are rearranged from an array of tandemly duplicated genes to form the antigen-binding portion of the Ig heavy (H) and light (L) chains. There is one H chain locus and 1-3 L chain isotypes, but only one H and L chain combination is used to form the receptor that is expressed on the B cell surface. Mechanisms involving not only hierarchical activation of the 3-4 loci but also feedback by a successful rearrangement are thought to restrict expression to one allele and one isotype, resulting in the expression of one receptor per cell. In higher vertebrates rearrangement thus not only creates a diverse antibody repertoire but plays a part in the regulation of immunoglobulin (Ig) gene expression. The basis for specificity is the clonal expression of receptors ; the basis for memory is the mitotic expansion of specific clones activated during the immune response.
In mammals clonal expression of the B cell receptor is suggested to result from several mechanisms: selection of one allele prior to rearrangement, favored recombination signal sequences (RSS), feedback inhibition whereby one allele generates a productive rearrangement which inhibits rearrangement of the other. These are all feasible within the confines of a system consisting of a few loci and where a hierarchy of activation is proposed -- H chain before L chain; kappa L chain before lambda L chain.
However, the regulatory role of rearrangement is one that evolved since its incipience in the ancestral jawed vertebrate. In representatives of the earliest vertebrates, the sharks and skates, there are hundreds of H and L chain loci, each consisting of a few gene segments that can recombine (VH-D-D-J-C and VL-J-C, respectively). Each locus functions independently. This is an early, alternative form of the antigen receptor gene organization (Figure 2). Another unusual feature of the shark Ig loci is that some contain partially or fully assembled VHD, VHDD, VHDDJ and VLJ sequences in the germline. Given these features, regulation of Ig expression in the shark cannot be easily explained by theories based on the assumptions made about rearrangement and regulation of Ig expression in mammals. In the shark, even if H chain is activated before L chain, there is not one but multiple H chain loci that can continue to rearrange. Among these H chain genes and within L isotypes the RSS are similar or identical; moreover, some of these genes are already rearranged. However, these germline-joined V genes do not pre-empt rearrangement at other loci and take over the system, indicating that the existence of rearranged sequence by itself is not sufficient. Furthermore, selective activation as a result of asynchronous replication of one chromosome is unlikely in shark, because our preliminary results show that the split and germline-joined genes are closely linked.
We ask: When 200 Ig loci are available, is clonal receptor expression still achieved? If not, to what extent is an antibody response affected; but if so, how is this regulated? These are questions fundamental to any system of multiple genes, like olfactory receptors, where function is integral to regulated expression.
What is the role of the germline-rearranged Ig genes when the recombination step is not needed? Although junctional diversity appears to be pre-empted in the germline-joined genes, they in fact do not remain invariant sequences. They (and rearranging Ig genes) undergo antigen-dependent hypermutation. We have established that not only does extensive somatic hypermutation occur at the shark Ig loci, but that the nature of the mutational mechanism(s) may be different. Our published and preliminary results show that half of somatic mutations are conventional point mutations but the other half consists of tandem changes of 2-5 bp. Any group of contiguous substitutions is initiated as a single event. This means that any "hit" changing two nucleotides in tandem will result in the replacement substitution of at least one amino acid. This is a remarkably efficient pathway of altering antibody specificities and the antibody response. We are in the process of characterizing this unusual mechanism and its relationship to the one in humans and mouse that produces only point mutations.
Since hypermutation occurs at the many H and L chain clusters in the shark, then the cis-acting elements regulating the phenomenon existed in the ancestral Ig gene, previous to H and L chain divergence. That is, before the formation of the 2H/2L chain basic unit that is present in all modern jawed vertebrates. Thus hypermutation is an ancient mechanism. The process for Ig diversification consists of two pathways (rearrangement, somatic hypermutation) involving DNA breaks. We know how old a mechanism rearrangement is - recombining Ig and TCR have only been found in jawed vertebrates but not in representatives of the earliest vertebrates, jawless fishes (hagfish, lamprey). Yet, Ig and TCR genes would have had to evolve from genes related to innate defense; the rearranging mechanism was selected for in evolution because it further diversified genes that already required sequence heterogeneity for their function. Perhaps these early (Figure 2, second line, pink box) non-rearranging immune genes hypermutated?
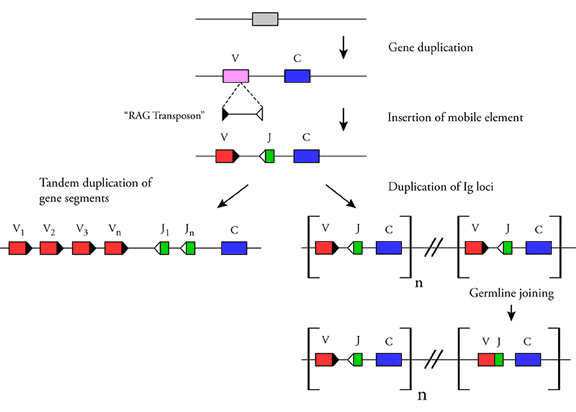
Figure 2. Hypothetical scheme for origin of antigen-receptor genes. The Ig superfamily is an ancient one, with members identifiable in sponges. A primordial Ig domain gave rise to V and C domains. The RAG transposon entered the lineage of the ancestral jawed vertebrate, and inserted the mobile element with RSS (indicated as triangles) into an archaic V domain, splitting it into two (V and J) gene segments. Thus, in (perhaps) one stroke, gene segments with the capability to recombine were created along with the installation of the transposase/recombinase RAG in the vertebrate genome. With the establishment of V(D)J recombination, two alternative antigen receptor gene organizations developed. Tandem duplication of the gene segments occurred at the one IgH or IgL locus in tetrapods (left). In elasmobranch IgH and IgL, duplication of the gene segments along with the C exons resulted in multiple loci (right). Rearrangement of V gene segments at some loci to VJ (IgL) and to VD, VDD, or VDDJ (IgH) occurred in the germ cells; many of the fully assembled V regions appear to be functional. (from "Origin of V(D)J Diversification", see bibliography). David Feliciano designed this figure.
Personnel
Karolina Malecek, Laboratory Manager
Victor Lee
Julie Brandman
Reviewer for various journals and granting agencies, including the National Science
Foundation and the National Institutes of Health
Editorial Advisory Board, Developmental and Comparative Immunology
Member, Expert Scientific Council, Protein Information Resource
- Lee, S.S., Fitch, D., Flajnik, M. F., and Hsu, E. (2000). Rearrangement of immunoglobulin genes in shark germ cells. J. Exp. Med. 191, 1637-1648. (See Commentary)
- Lee, S. S., Tranchina, D. S., Ohta, Y., Flajnik, M. F., and Hsu, E. (2002). Hypermutation in shark immunoglobulin light chain genes results in contiguous substitutions. Immunity 16, 571-582.
- Malecek, K., Lee, V., Feng, W., Huang, J.L., Flajnik, M.F., Ohta, Y., and Hsu, E. (2008) Immunoglobulin heavy chain exclusion in the shark. PLoS Biol 6: e157.
- Hsu, E. (2011). The invention of lymphocytes. Curr. Opin. Immunol. 23, 1-7.
- Zhu, C., Lee, V., Finn, A., Senger, K., Zarrin, A.A., Du Pasquier, L., and Hsu, E. (2012). Origin of immunoglobulin isotype switching. Curr. Biol. 22, 872-880.
- Hsu, E., and Lewis, S.M. (2015) The origin of V(D)J diversification. In: Molecular Biology of B cells, Alt, F.W., Honjo, T., Radbruch, A., and Reth, M., eds. Elsevier, Academic Press, Amsterdam, pp. 133-148.
- Hsu, E. (2016) Assembly and expression of shark immunoglobulin genes. J. Immunol. 196: 3517-3523.
- Iacoangeli, A., Lui, A., Haines, A., Ota, Y., Flajnik, M. F., and Hsu, E. (2017). Evidence for immunoglobulin light chain isotype exclusion in shark B lymphocytes suggests ordered mechanisms. J. Immunol. 199, 1875-1885.
- Anumukonda, K., Francis, M., Currie, P., Tulenko, F., and Hsu, E. (2022). Heavy chain-only antibody genes in fish evolved to generate unique CDR3 repertoire. Eur. J. Immunol. 52, 247-260. Highlighted in "In This Issue", Eur. J. Immunol. 52, 187.
- Hsu, E. (2024). The immune system and immunity in cartilaginous fish. In: The Encyclopedia of Immunobiology, 2nd edition. Reference Module in Life Sciences. Elsevier Ltd. Available on request.